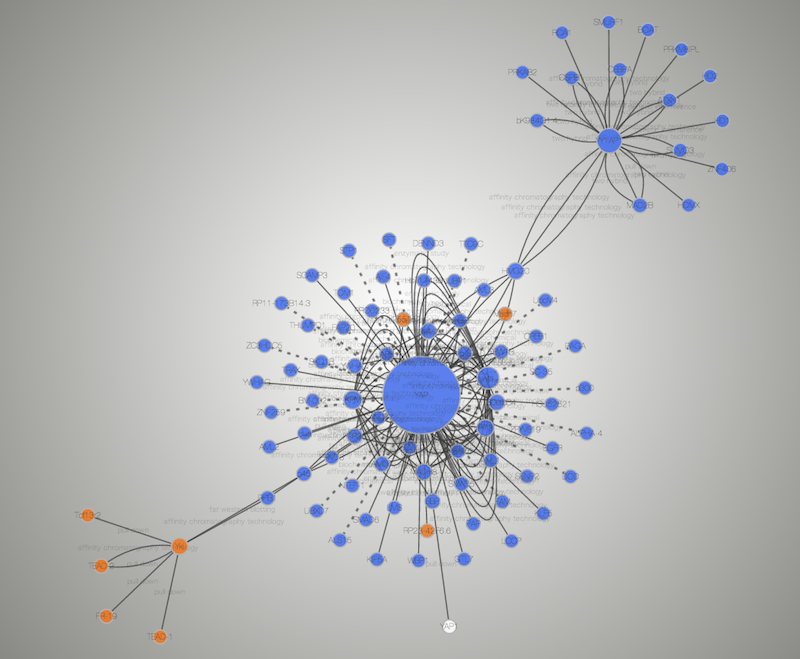

The availability of high-throughput molecular interaction network provides and the application of network analysis tools such as clustering or graph partitioning have proved valuable in disease gene prioritization. Genome-wide association studies and linkage analysis have been pivotal in the identification of candidate genes, however, the large list of resultant genes returned are time-consuming and expensive to analyze. Identification of candidate genes associated with physiological disorders are a fundamental task in the analysis of complex diseases. Research by discovered that genes connected to diseases with similar phenotypes are more likely to interact directly with each other. demonstrated that the majority of disease genes are nonessential and located in the periphery of functional networks. have indicated that the alterations in the physical interaction network may be a indicator of breast cancer prognosis. 
analyzed topological features of a PPIN and observed that hereditary disease-genes from the Online Mendelian Inheritance in Man (OMIM) database have a larger degree and tendency to interact with other disease-genes in literature curated networks. Network theory is making important contributions to the topological study of biological networks, such as Protein-Protein Interaction Networks (PPIN). Via the analysis of network topology and dynamics, key discoveries have been made including identification of novel disease genes and pathways, biomarkers and drug targets for disease. The emergence of network medicine has explored disease complexity through the systematic identification of disease pathways and modules. The rapid accumulation of high-throughput data along with advances in network biology have been fundamental in improving our knowledge of biological systems and complex disease. This work provides a foundation for future investigation of diverse heterogeneous data integration for disease-gene prioritization. Relatively high predictive performance (AUC: 0.70) was observed when classifying AD and normal gene expression profiles from individuals using leave-one-out cross validation. Biological process enrichment analysis revealed the prioritized genes are modulated in AD pathogenesis including: regulation of neurogenesis and generation of neurons. This approach correctly identified key AD susceptible genes: PSEN1 and TRAF1. The pipeline was applied to prioritize Alzheimer's Disease (AD) genes, whereby a list of 32 prioritized genes was generated. Diverse heterogeneous data including: gene-expression, protein-protein interaction network, ontology-based similarity and topological measures and tissue-specific are integrated. In this paper we propose a computational pipeline for the prioritization of disease-gene candidates. The integration of protein-protein interaction networks along with disease datasets and contextual information is an important tool in unraveling the molecular basis of diseases. The application of gene prioritization can enhance our understanding of disease mechanisms and aid in the discovery of drug targets. 
An important challenge now is to identify meaningful disease-associated genes from a long list of candidate genes implicated by these analyses. Genome-wide linkage and association studies have made advancements in identifying genetic variants that underpin human disease. The identification of genes and uncovering the role they play in diseases is an important and complex challenge.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
